Millard Group - Molecular mechanisms for wiring the brain
The overall goal of the Millard lab is to understand how specificity is generated in the brain.
This problem is best exemplified by considering that 100 trillion synapses are generated and maintained in the human brain using a toolkit of only 20,000 genes. We have been approaching this problem using molecular genetics in the fruit fly, Drosophila melanogaster. Most projects in the lab revolve around how a broadly expressed cell surface protein, called Down syndrome cell adhesion molecule 2 (Dscam2), is able to perform specific functions in different neurons. We are also interested in mechanisms of neurological disease, particularly those that involve changes in synaptic function.
Projects in the lab
 We use Drosophila molecular genetics to understand:
We use Drosophila molecular genetics to understand:
- how brain wiring proteins work at the molecular level.
- how brain structures and neural circuits form.
- how the brain generates sufficient protein diversity to specify its numerous synaptic connections - focus on alternative splicing.
- how cell surface proteins regulate synaptic physiology.
We also use flies to model human disease processes. ALS is our current focus.
- Modelling neurodegeneration in the fly eye using candidate ALS risk factors.
- Studying how ALS risk factors affect motor neuron morphology, synaptic makeup and physiology.
Lab work flow and techniques
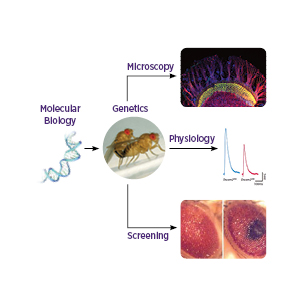
We use molecular biology to manipulate genes or the genome, perform genetics to generate animals of the appropriate genotype and then score the resulting phenotypes using confocal microscopy, electrophysiology or external phenotypes in the fly.
- CRISPR
- RNAi
- Cell-specific overexpression
- Mosaic experiments (single cells, rather than the entire organism, manipulated)
- High resolution imaging
- High-throughput screening in a whole animal model
We are always interested in recruiting enthusiastic scientists and welcome new ideas and techniques. Please contact Sean to enquire about Honours, Masters, PhD or Postdoctoral positions in the lab. Below is a list of ongoing projects.
Group Head
Staff
Students
Former students
- Sarah Kerwin (PhD) – Postdoctoral Fellow at Rockefeller University (Bargmann lab)
- Amber Kewin (MPhil) – Research Assistant at the Queensland Brain Institute (van Swinderen lab)
- Lorenzo Odierna (PhD) – Postdoctoral Fellow at the University of Tasmania (Blizzard lab)
- Joshua Li (PhD) – Postdoctoral Fellow at Harvard University (Perrimon lab)
- Kevin Mutemi (MPhil) – PhD student at EMBL Heidelberg (Arendt lab)
For a full list of publications, visit eSpace
2020
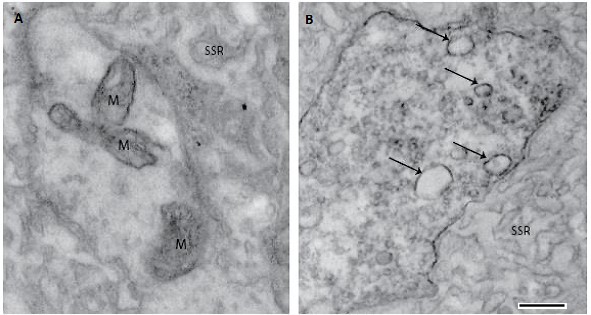
Dscam2 suppresses synaptic strength through a PI3K-dependent endosomal pathway.
Odierna GL, Kerwin SK, Harris LE, Shin GJ, Lavidis NA, Noakes PG, Millard SS.
J Cell Biol. 2020 Jun 1;219(6):e201909143. doi: 10.1083/jcb.201909143.
PMID: 32259198
Elevated levels of Drosophila Wdr62 promote glial cell growth and proliferation through AURKA signalling to AKT and MYC.
Shohayeb B, Mitchell N, Millard SS, Quinn LM, Ng DCH.
Biochim Biophys Acta Mol Cell Res. 2020 Jul;1867(7):118713. doi: 10.1016/j.bbamcr.2020.118713. Epub 2020 Apr 1.
PMID: 32246948
The association of microcephaly protein WDR62 with CPAP/IFT88 is required for cilia formation and neocortical development.
Shohayeb B, Ho U, Yeap YY, Parton RG, Millard SS, Xu Z, Piper M, Ng DCH.
Hum Mol Genet. 2020 Jan 15;29(2):248-263. doi: 10.1093/hmg/ddz281.
PMID: 31816041
2019
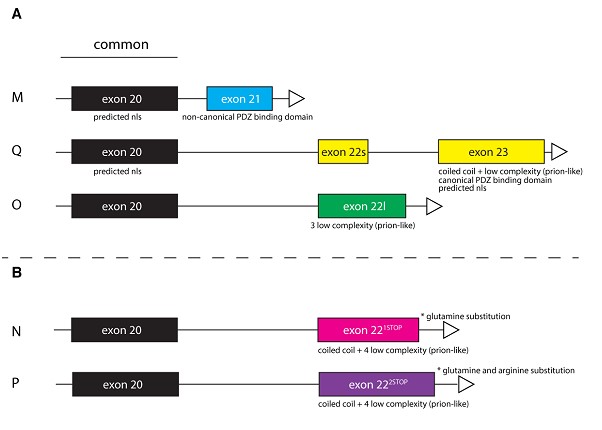
Li, JS and Millard SS (2019). Deterministic splicing of Dscam2 is regulated by Muscleblind. Sci Adv 5(1): eaav1678.
2018
Millard SS, Pecot MY (2018). Strategies for assembling columns and layers in the Drosophila visual system. Neural Development 13(1):11.
Kerwin SK, Li JS, Noakes PG, Shin GJ, Millard SS (2018). Regulated alternative splicing of Drosophila Dscam2 is necessary for attaining the appropriate number of photoreceptor synapses. Genetics 208 (2): 717-28.
2017
Lim NR, Shohayeb B, Zaytseva O, Mitchell N, Millard SS, Ng DCH, and Quinn LM (2017). Glial-specific functions of microcephaly protein WDR62 and interaction with the mitotic kinase AURKA are essential for Drosophila brain growth. Stem Cell Reports 9: 1-10.
2016
Tadros W, Xu S, Akin O, Yi CH, Shin GJ, Millard SS and Zipursky SL (2016). Dscam proteins direct dendritic targeting through adhesion. Neuron 89 (3):480-93.
2015
Li JS, Shin GJ, Millard SS (2015). Neuronal cell-type-specific alternative splicing: A mechanism for specifying connections in the brain? Neurogenesis 2:1, e1122699: 1-5.
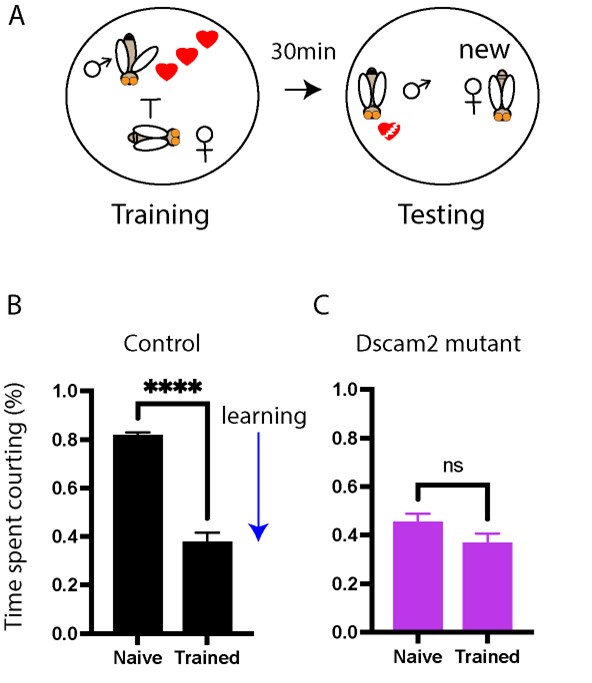
Bosch DS, van Swinderen B and Millard SS (2015). Dscam2 affects visual perception in Drosophila melanogaster. Front. Behav. Neurosci. 9:149. doi: 10.3389/fnbeh.2015.00149.
2014
Lah GJ, Li JS, Millard SS (2014). Cell-specific alternative splicing of Drosophila Dscam2 is crucial for proper neuronal wiring. Neuron 17;83(6):1376-88.
2013
Li Q, Ha TS, Okuwa S, Wang Y, Wang Q Millard SS, Smith DP and Volkan-Cayirlioglu P (2013). Combinatorial rules of precursor specification underlying olfactory neuron diversity. Current Biol. 23, 2481-2490.
Paulk A, Millard SS, van Swinderen B. (2013). Vision in Drosophila: Seeing the World Through a Model's Eyes. Ann. Rev. Entomol. 58:313-32.
2010
Millard SS, Lu Z, Zipursky SL, Meinertzhagen IA. (2010) Drosophila Dscam proteins regulate postsynaptic specificity at multiple-contact synapses. Neuron 9;67(5):761-8.
2008
Millard SS, Zipursky SL. (2008) Dscam-mediated Repulsion Controls Tiling and Self-avoidance. Current Opinion in Neurobiology 18(1):84-9.
Hattori D, Millard SS, Wojtowicz W, Zipursky SL. (2008) Dscam-mediated Cell Recognition Regulates Neural Circuit Formation. Annual Reviews in Cell and Developmental Biology 24:597-620.
2007
Millard SS, Flanagan JJ, Pappu KS, Wu W, Zipursky SL. (2007) Dscam2 mediates axonal tiling in the Drosophila visual system. Nature 447(7145):720-4.
Find out more about our diverse range of research interests.